The exceptions are our daily pain but the exceptions hard to explain are more than that. I faced one of them one day when I was integrating Apache Spark SQL on EMR.
What would it take for you to trust your Databricks pipelines in production?
A 3-day bug hunt on a 3-person team costs up to €7,200 in lost engineering time. This workshop teaches you to prevent that — unit tests, data tests, and integration tests for PySpark and Databricks Lakeflow, including Spark Declarative Pipelines.

Konieczny
The post is composed of 2 sections. The first one introduces the problem. The second one shows some internals leading to it. Spoiler: I don't have a solution for that. But I hope that the internal details will help you to understand why this error happens.
FileAlreadyExistsException with overwrite mode
Whenever I can, I try to adopt the principle of immutable tables proposed by Maxime Beauchemins in his Functional Data Engineering — a modern paradigm for batch data processing post. In Apache Spark you can achieve it quite easily with SaveMode.Overwrite that will simply drop all existing data in the sink and replace it with the new generated one. The code that I will analyze looks like that:
object FileAlreadyExistProblem {
import org.apache.spark.sql.{SaveMode, SparkSession}
def main(args: Array[String]): Unit = {
val sparkSession: SparkSession = SparkSession.builder()
.appName("Spark SQL FileAlreadyExist error example")
.getOrCreate()
import sparkSession.implicits._
val dataset = Seq(
(1, "a"), (2, "b"), (3, "c"), (4, "d"), (0, ".")
).toDF("nr", "letter").repartition(3)
val datasetWithDivisionResults = dataset.map(row => {
try {
30/row.getAs[Int]("nr")
} catch {
case e => {
val previousFlag = FailedFlag.isFailed
FailedFlag.isFailed = true
if (!previousFlag) throw e else 0
}
}
})
datasetWithDivisionResults
.write.mode(SaveMode.Overwrite)
.json("s3n://my-test-bucket/test-file-already-exists")
}
}
object FailedFlag {
var isFailed = false
}
As you can see, the operation is quite simple. We're taking some data from an in-memory relationship, map it into another format to finally write to some bucket on S3. An important thing to notice is that the operation fails during the first mapping execution for the column with 0. After executing that code on EMR (eg. spark-submit --deploy-mode cluster --master yarn --class com.waitingforcode.FileAlreadyExistProblem s3://my-jar-bucket/test-program-jar-with-dependencies.jar), you should get this error:
19/09/19 20:19:49 ERROR Executor: Exception in task 0.1 in stage 1.0 (TID 8)
org.apache.hadoop.fs.FileAlreadyExistsException: File already exists: s3://my-test-bucket/test-file-already-exists/part-00000-005ee88c-e62d-47b4-b56c-b725db3cc9b7-c000.json
at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.RegularUploadPlanner.checkExistenceIfNotOverwriting(RegularUploadPlanner.java:36)
at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.RegularUploadPlanner.plan(RegularUploadPlanner.java:30)
at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.UploadPlannerChain.plan(UploadPlannerChain.java:37)
at com.amazon.ws.emr.hadoop.fs.s3n.S3NativeFileSystem.create(S3NativeFileSystem.java:601)
at org.apache.hadoop.fs.FileSystem.create(FileSystem.java:932)
at org.apache.hadoop.fs.FileSystem.create(FileSystem.java:913)
at org.apache.hadoop.fs.FileSystem.create(FileSystem.java:810)
at com.amazon.ws.emr.hadoop.fs.EmrFileSystem.create(EmrFileSystem.java:217)
at org.apache.spark.sql.execution.datasources.CodecStreams$.createOutputStream(CodecStreams.scala:81)
at org.apache.spark.sql.execution.datasources.CodecStreams$.createOutputStreamWriter(CodecStreams.scala:92)
at org.apache.spark.sql.execution.datasources.json.JsonOutputWriter.(JsonFileFormat.scala:183)
at org.apache.spark.sql.execution.datasources.json.JsonFileFormat$$anon$1.newInstance(JsonFileFormat.scala:82)
at org.apache.spark.sql.execution.datasources.SingleDirectoryDataWriter.newOutputWriter(FileFormatDataWriter.scala:120)
at org.apache.spark.sql.execution.datasources.SingleDirectoryDataWriter.(FileFormatDataWriter.scala:108)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$.org$apache$spark$sql$execution$datasources$FileFormatWriter$$executeTask(FileFormatWriter.scala:236)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$$anonfun$write$1.apply(FileFormatWriter.scala:170)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$$anonfun$write$1.apply(FileFormatWriter.scala:169)
at org.apache.spark.scheduler.ResultTask.runTask(ResultTask.scala:90)
at org.apache.spark.scheduler.Task.run(Task.scala:121)
at org.apache.spark.executor.Executor$TaskRunner$$anonfun$10.apply(Executor.scala:408)
at org.apache.spark.util.Utils$.tryWithSafeFinally(Utils.scala:1360)
at org.apache.spark.executor.Executor$TaskRunner.run(Executor.scala:414)
at java.util.concurrent.ThreadPoolExecutor.runWorker(ThreadPoolExecutor.java:1149)
at java.util.concurrent.ThreadPoolExecutor$Worker.run(ThreadPoolExecutor.java:624)
at java.lang.Thread.run(Thread.java:748)
Of course, the real error of the above exception was:
19/09/19 20:25:52 ERROR Utils: Aborting task
java.lang.ArithmeticException: / by zero
at com.waitingforcode.FileAlreadyExistProblem$$anonfun$1.apply(FileAlreadyExistProblem.scala:23)
at com.waitingforcode.FileAlreadyExistProblem$$anonfun$1.apply(FileAlreadyExistProblem.scala:21)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage1.mapelements_doConsume_0$(Unknown Source)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage1.deserializetoobject_doConsume_0$(Unknown Source)
at org.apache.spark.sql.catalyst.expressions.GeneratedClass$GeneratedIteratorForCodegenStage1.processNext(Unknown Source)
at org.apache.spark.sql.execution.BufferedRowIterator.hasNext(BufferedRowIterator.java:43)
at org.apache.spark.sql.execution.WholeStageCodegenExec$$anonfun$13$$anon$1.hasNext(WholeStageCodegenExec.scala:636)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$$anonfun$org$apache$spark$sql$execution$datasources$FileFormatWriter$$executeTask$3.apply(FileFormatWriter.scala:244)
at org.apache.spark.sql.execution.datasources.FileFormatWriter$$anonfun$org$apache$spark$sql$execution$datasources$FileFormatWriter$$executeTask$3.apply(FileFormatWriter.scala:242)
at org.apache.spark.util.Utils$.tryWithSafeFinallyAndFailureCallbacks(Utils.scala:1394)
Stack trace analysis
Since this behavior is surprising - after all I want to overwrite everything - I started to investigate. Unfortunately, I didn't find the code source of AWS EMR classes (UploadPlannerChain, RegularUploadPlanner), so I had to stop at one level before.
I started my analysis by JsonOutputWriter and its line 183:
private val writer = CodecStreams.createOutputStreamWriter(
context, new Path(path), encoding)
As you can see here, it creates an output stream writer that will be used later to write one row at a time from this method:
override def write(row: InternalRow): Unit = {
gen.write(row)
gen.writeLineEnding()
}
But since the stack trace points to the CodecsStream class, I took a look at it as well. And what I saw is probably the problem (line 81, so just before moving to com.amazon.ws.emr classes):
def createOutputStream(context: JobContext, file: Path): OutputStream = {
val fs = file.getFileSystem(context.getConfiguration)
val outputStream: OutputStream = fs.create(file, false)
A hint? The invoked create method looks like that:
/**
* Create an FSDataOutputStream at the indicated Path.
* @param f the file to create
* @param overwrite if a file with this name already exists, then if true,
* the file will be overwritten, and if false an exception will be thrown.
*/
public FSDataOutputStream create(Path f, boolean overwrite)
throws IOException {
return create(f, overwrite,
getConf().getInt("io.file.buffer.size", 4096),
getDefaultReplication(f),
getDefaultBlockSize(f));
}
As you can see, the false value is a flag to say whether the writing should overwrite already existent file. And since it's hardcoded, we can't change it. Below you can find the bucket after the failure:
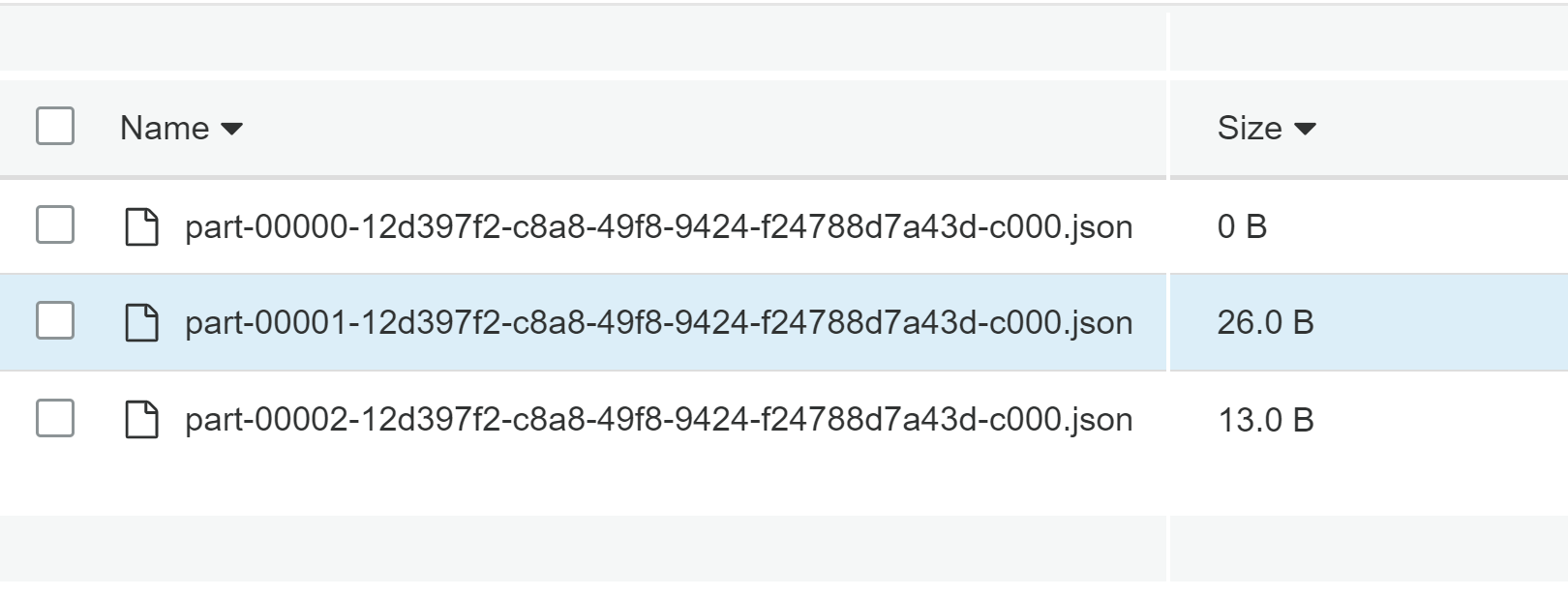
The overwrite value hardcoded to false makes sense with the rest of the stack trace:
at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.RegularUploadPlanner .checkExistenceIfNotOverwriting(RegularUploadPlanner.java:36) at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.RegularUploadPlanner .plan(RegularUploadPlanner.java:30) at com.amazon.ws.emr.hadoop.fs.s3.upload.plan.UploadPlannerChain .plan(UploadPlannerChain.java:37)
As you can see, the method called after creating not overwritable stream is checkExistenceIfNotOverwriting. Regarding the name of this function, we can suppose that it will make the processing fail if the file already exists. I tried to find other methods of RegularUploadPlanner to get a better idea about potential execution but without success.
In this post you can discover that save mode doesn't apply to the whole execution. In the case of a task retry, the overwrite flag is hardcoded to false which can make the task fail in case of multiple executions, as illustrated in this article. If you have any clue how to solve it, your comment is welcome. You'll help not only me but also other people since the question about FileAlreadyExistsException was asked twice (links in Read also section).
Data Engineering Design Patterns

Looking for a book that defines and solves most common data engineering problems? I wrote
one on that topic! You can read it online
on the O'Reilly platform,
or get a print copy on Amazon.
I also help solve your data engineering problems contact@waitingforcode.com 📩
Read also about FileAlreadyExistsException at task retry on EMR here:
- File already exists error writing new files from dataframe Spark FileAlreadyExistsException on Stage Failure
Related blog posts:
- Clean architecture for PySpark
- ASOF Join in Apache Spark SQL
- Outer operations in Apache Spark, or why I consider NULLs as NullPointerException
Just before publishing my Spark+AI Summit follow-up posts, I'd like to share with you an investigation I made to understand a FileAlreadyExistsException when running #ApacheSpark on #AWS #EMR https://t.co/KD53i7RiOj
— Bartosz Konieczny (@waitingforcode) October 19, 2019

