After last week's global overview of graph representation in GraphX module, it's time to go a little bit deeper and analyze the 2 main components of graphs: vertices and edges. We'll begin here with the former ones.
What would it take for you to trust your Databricks pipelines in production?
A 3-day bug hunt on a 3-person team costs up to €7,200 in lost engineering time. This workshop teaches you to prevent that — unit tests, data tests, and integration tests for PySpark and Databricks Lakeflow, including Spark Declarative Pipelines.

Konieczny
In this post we'll focus on VertexRDD which is GraphX's representation of vertices. Its first section presents general points about it and shows some basic manipulations. The next part is dedicated to the routing table data structure. The last one talks about another component of VertexRDD which is the bitmask.
VertexRDD
Graph object in GraphX is composed of 2 RDDs: VertexRDD and EdgeRDD. Thanks to this separation we can apply different transformations to each of them. VertexRDD supports all manipulations known from RDD, such as filtering or mapping:
it should "map and filter the vertices" in {
val vertices = TestSparkContext.parallelize(
Seq((1L, "A"), (2L, "B"), (3L, "C"))
)
val edges = TestSparkContext.parallelize(
Seq(Edge(1L, 2L, ""))
)
val graph = Graph(vertices, edges)
val mappedAndFilteredVertices = graph.vertices.map(idWithLetter => s"${idWithLetter._2}${idWithLetter._2}")
.filter(letter => letter != "CC")
val collectedAttributes = mappedAndFilteredVertices.collect()
collectedAttributes should have size 2
collectedAttributes should contain allOf("AA", "BB")
}
In addition to them, VertexRDD has also some more graph-specific operations, such as aggregateUsingIndex to merge vertices of the same id in 2 different datasets, innerJoin to join 2 different datasets of vertices, or diff to compare 2 graphs:
it should "join 2 vertices datasets and merge their attributes" in {
val sharedVertices = Seq((1L, "A"), (2L, "B"), (3L, "C"))
val vertices = TestSparkContext.parallelize(sharedVertices)
val edges = TestSparkContext.parallelize(Seq(Edge(1L, 2L, "")))
val graph1 = Graph(vertices, edges)
val vertices2 = TestSparkContext.parallelize(sharedVertices ++ Seq((4L, "D")))
val graph2 = Graph(vertices2, edges)
val joinedVertices = graph1.vertices.innerJoin(graph2.vertices) {
case (vertexId, vertex1Attr, vertex2Attr) => s"${vertex1Attr}${vertex2Attr}"
}
val collectedVertices = joinedVertices.collect()
collectedVertices should have size 3
collectedVertices should contain allOf((1L, "AA"), (2L, "BB"), (3L, "CC"))
}
it should "get vertices with different values in the second dataset" in {
val sharedVertices = Seq((1L, "A"), (2L, "B"), (3L, "C"))
val vertices = TestSparkContext.parallelize(sharedVertices ++ Seq((4L, "D"), (5L, "E")))
val edges = TestSparkContext.parallelize(Seq(Edge(1L, 2L, "")))
val graph1 = Graph(vertices, edges)
val vertices2 = TestSparkContext.parallelize(sharedVertices ++ Seq((4L, "DD"), (5L, "EE"), (6L, "FF")))
val graph2 = Graph(vertices2, edges)
val joinedVertices = graph1.vertices.diff(graph2.vertices)
val collectedVertices = joinedVertices.collect()
collectedVertices should have size 2
// As you can see, it doesn't return the vertices of the graph2 that are not in the graph1
collectedVertices should contain allOf((4L, "DD"), (5L, "EE"))
}
Internally VertexRDD is a children class for RDD[(VertexId, VD)] where the former type represents the ids of vertices and the latter their attributes. Even though the id is represented here as a VertexId type, it's in fact an alias for a Long type. The id is used by org.apache.spark.HashPartitioner to define the partition storing given vertex.
Aside of VertexId and attributes, each VertexRDD partition has also 2 other properties: routing table and bitset. I'll detail both in the next 2 sections.
VertexRDD and routing table
In the previous post about graphs representation in Apache Spark GraphX, you could see the use of triplets which are the structures composed of vertices, their attributes and the edge between them. As told, using them involves moving vertices to the edges partitions. We can notice that in org.apache.spark.graphx.impl.ReplicatedVertexView#upgrade(vertices: VertexRDD[VD], includeSrc: Boolean, includeDst: Boolean) method:
val shippedVerts: RDD[(Int, VertexAttributeBlock[VD])] =
vertices.shipVertexAttributes(shipSrc, shipDst)
.setName("ReplicatedVertexView.upgrade(%s, %s) - shippedVerts %s %s (broadcast)".format(
includeSrc, includeDst, shipSrc, shipDst))
.partitionBy(edges.partitioner.get)
val newEdges = edges.withPartitionsRDD(edges.partitionsRDD.zipPartitions(shippedVerts) {
(ePartIter, shippedVertsIter) => ePartIter.map {
case (pid, edgePartition) =>
(pid, edgePartition.updateVertices(shippedVertsIter.flatMap(_._2.iterator)))
}
})
The logic of shipping the attributes to the edges partitions is defined in ShippableVertexPartition's shipVertexAttributes(shipSrc: Boolean, shipDst: Boolean):
Iterator.tabulate(routingTable.numEdgePartitions) { pid =>
routingTable.foreachWithinEdgePartition(pid, shipSrc, shipDst) { vid =>
if (isDefined(vid)) {
vids += vid
attrs += this(vid)
}
i += 1
}
(pid, new VertexAttributeBlock(vids.trim().array, attrs.trim().array))
It's there where the aspect of routing table appears for the first time. As you can see in the snippet, we iterate over all edge partitions related to the current partition's vertices and collect the attributes sequentially. In the end, we construct a block and send everything to edges partitions. The code is self-explanatory and shows pretty clearly what the routing table is.
In fact, the GraphX supposes that in most of the cases, the number of edges will be much bigger than the number of vertices. Therefore, during the triplets construction, the library ships vertices attributes to the edges and not inversely. The routing table is then a local table for each vertex partition storing the edge partitions where all partition's vertices are present. In the image below we can see that the node A is defined in all edge partitions and the other vertices in only one:
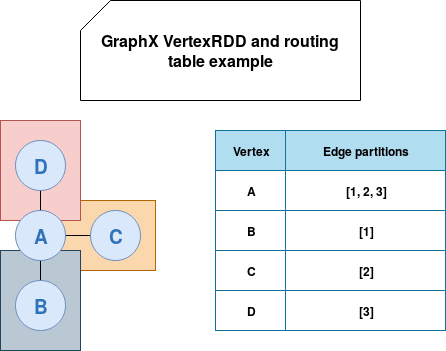
Bitmask
In addition to the routing table, each vertex partition has also its own bitmask table. The role of this structure is less obvious but also pretty useful in the processing of VertexRDD. In the code we can observe its use in org.apache.spark.graphx.impl.VertexPartitionBaseOps#withMask(org.apache.spark.graphx.impl.VertexPartitionBaseOps#withMask) method whose implementation depends on the vertex partition type. The bitmask is used in already presented map and filter operations:
def map[VD2: ClassTag](f: (VertexId, VD) => VD2): Self[VD2] = {
// Construct a view of the map transformation
val newValues = new Array[VD2](self.capacity)
var i = self.mask.nextSetBit(0)
while (i >= 0) {
newValues(i) = f(self.index.getValue(i), self.values(i))
i = self.mask.nextSetBit(i + 1)
}
this.withValues(newValues)
}
def filter(pred: (VertexId, VD) => Boolean): Self[VD] = {
// Allocate the array to store the results into
val newMask = new BitSet(self.capacity)
// Iterate over the active bits in the old mask and evaluate the predicate
var i = self.mask.nextSetBit(0)
while (i >= 0) {
if (pred(self.index.getValue(i), self.values(i))) {
newMask.set(i)
}
i = self.mask.nextSetBit(i + 1)
}
this.withMask(newMask)
}
The code above shows one of the already mentioned properties of GrapX - the library tries to reuse the most of attributes possible. We can see this in the filter method where we don't copy any values. Instead, we only keep the vertices matching the predicate in the bitmask. Thanks to that in the map we'll have much fewer data to process.
Filter and map are not the single operations using the bitmask. We can notice the real power of bitmasks in the operations combining different graphs, like diff or innerJoin which compute the bit-wise AND. Just to recall, the AND returns 1 only if the bits at the same position in 2 different bitsets are equal to 1. After applying that operation we can simply iterate over so combined bitmask and use our function computing the value after join:
val newMask = self.mask & other.mask
val newValues = new Array[VD2](self.capacity)
var i = newMask.nextSetBit(0)
while (i >= 0) {
newValues(i) = f(self.index.getValue(i), self.values(i), other.values(i))
i = newMask.nextSetBit(i + 1)
}
this.withValues(newValues).withMask(newMask)
VertexRDD is one of the components of Graph representation. It shares a lot of characteristics with other RDDs, such as mapping and filtering transformations, partitioning or caching. However, it also brings its own graph-specific features. We've discovered 2 of them: routing table and bitmask. The former is used to efficiently dispatch vertex attributes to the edge partitions where the given vertex is present. The bitmask is used to hide the vertices not meeting the predicate (filter) or to efficiently join 2 vertices datasets.
Data Engineering Design Patterns

Looking for a book that defines and solves most common data engineering problems? I wrote
one on that topic! You can read it online
on the O'Reilly platform,
or get a print copy on Amazon.
I also help solve your data engineering problems contact@waitingforcode.com 📩
Related blog posts:
- Edge representation in Apache Spark GraphX
- Graphs representation in Apache Spark GraphX
- PCollection - data representation in Apache Beam
- Dataset in Spark SQL
- DStream in Spark Streaming
#ApacheSpark #GraphX graph abstraction is composed of 2 RDDs. One of them represents the vertices and today's post https://t.co/DpN5201Fve covers its main aspects as for instance routing tables.
— Bartosz Konieczny (@waitingforcode) December 19, 2018

