It's always a huge pleasure to see the PySpark API covering more and more Scala API features. Starting from Apache Spark 3.4.0 you can even write arbitrary stateful processing jobs! But since the API is a little bit different than the one available on the Scala side, I wanted to take a deeper look.
What would it take for you to trust your Databricks pipelines in production?
A 3-day bug hunt on a 3-person team costs up to €7,200 in lost engineering time. This workshop teaches you to prevent that — unit tests, data tests, and integration tests for PySpark and Databricks Lakeflow, including Spark Declarative Pipelines.

Konieczny
The applyInPandasWithState has several differences with its Scala counterpart that do make sense from the engineering standpoint.
- The function signature is heavier because it requires specifying the state schema and the output schema. Remember, Python is not strongly typed as Scala. These explicit definitions are here to bring some type safety to the transformation.
- The stateful mapping function - the one called under-the-hood by the applyInPandasWithState - operates on Iterable[pandas.DataFrame]. Here too, using Pandas makes sense because of its underlying optimizations, including Apache Arrow batching. Additionally, Pandas are also the input of the applyInPandas and having the same convention is more user-friendly.
- The stateful mapping function works on an Iterable and not on a pandas.DataFrame, you can read in the previous point. Indeed. The reason for that was to address the limitations of the applyInPandas scalability. One of its requirements was the data that must fit into memory. For the stateful version, it's addressed with a generator for the input and output parameters. That way, a big group dataset can be splitted into multiple RecordBatches and delivered iteratively without compromising the memory. Of course, if you decide to materialize that batch of records, the lazy loading won't help you but at least it guarantees that PySpark itself won't be the OOM reason.
- The state parameter of the stateful function is not your grouping key. Well, only a part of it is the grouping key. The other part is the type of the key expressed as a NumPy data type.
Implementation
After reading the list you may be thinking the feature is a copy-paste of the applyInPandas. Well, it has some similarities but in the end, Jungtaek Lim, the author of the feature, had to made many changes to plug the stateful part on the Scala API to the stateful part on the PySpark side. And before I explain them all to you, let me introduce the interactions diagram:
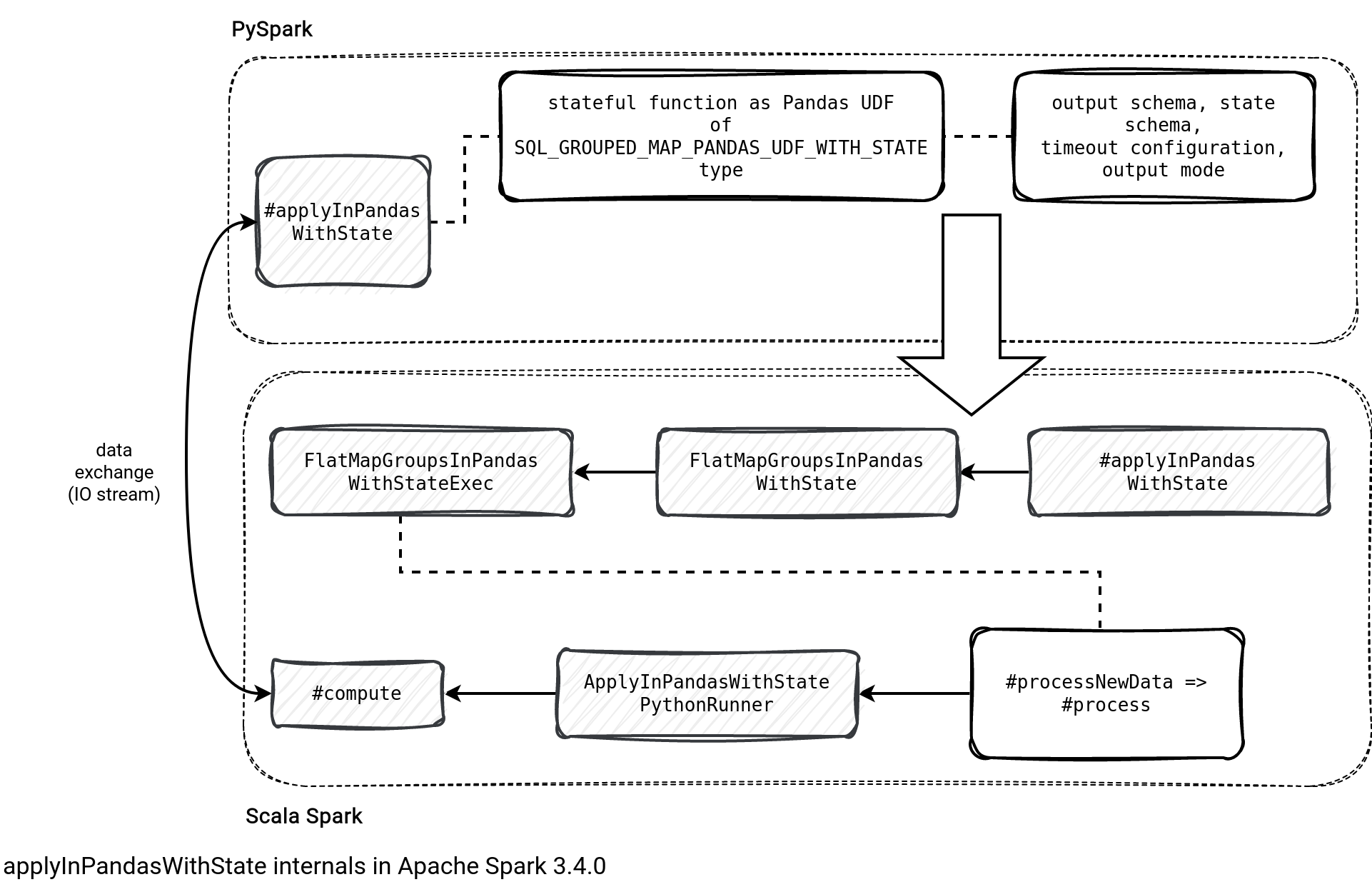
Everything starts in your PySpark job when you declare your stateful function with all the required parameters, including both schemas (output, state), output mode, and the timeout configuration. All is later transformed to a Python UDF (aka vectorized UDF) and passed to the JVM's applyInPandasWithState:
def applyInPandasWithState(
self, func: "PandasGroupedMapFunctionWithState",
outputStructType: Union[StructType, str],
stateStructType: Union[StructType, str],
outputMode: str, timeoutConf: str,
) -> DataFrame:
# ...
udf = pandas_udf(
func, # type: ignore[call-overload]
returnType=outputStructType,
functionType=PythonEvalType.SQL_GROUPED_MAP_PANDAS_UDF_WITH_STATE,
)
df = self._df
udf_column = udf(*[df[col] for col in df.columns])
jdf = self._jgd.applyInPandasWithState(
udf_column._jc.expr(),
self.session._jsparkSession.parseDataType(outputStructType.json()),
self.session._jsparkSession.parseDataType(stateStructType.json()),
outputMode,
timeoutConf,
)
return DataFrame(jdf, self.session)
At that moment the JVM side only knows about the function to run. It then creates the logical node (FlatMapGroupsInPandasWithState to hold the passed parameters alongside the function itself of the PythonUDF type. During the physical planning, the node gets resolved to FlatMapGroupsInPandasWithStateExec physical node that triggers data communication between the JVM and PVM in the process method.
The communication consists of creating a ApplyInPandasWithStatePythonRunner. The runner passes Pandas groups to the Python stateful function and reads back the function outcome as a tuple of state updates and generated output.
The state updates are required because the state management is still performed on the JVM side.
Tips, tricks, lessons learned
The first surprising thing - but I'm a JVM guy - was the output data definition. The Pandas initialization was failing with a ValueError: If using all scalar values, you must pass an index. It's a normal behavior if you pass a simple dict to the constructor.
The second point is about the grouping key. It's also valid for the Scala API but the impact may be more visible in the PySpark version due to this JVM <=> PVM communication. Your state doesn't need to store the grouping key because first, it's already stored as the key in the state store. Second, it's always passed to your stateful mapping function, so having it in the state structure adds these few extra bytes in the serialization loop.
Pandas transformations... If you have been writing your code with Scala API and have now to implement the stateful mapping in PySpark, you'll rely a lot on the Pandas transformation. They're mostly column-based and may appear to be unnatural in the beginning. Hopefully, after first tries-and-fails, you should switch between these 2 approaches easier.
Pandas transformations and interactivity. Being unfamiliar with Pandas I counted a lot on Python REPL and transformations on small and simple datasets for easier debugging.
When it comes to the state, you can read it but it won't be accessible as an easy-to-use class. Instead, it's returned as a tuple, for example start_time, end_time, = current_state.get. You need to be careful about the order of fields.
For a more low-level perspective, JVM passes Pandas data as one or multiple Apache Arrow batches. You shouldn't assume each group will receive only 1 batch of input records to process and that's the reason you'll often find for loops the stateful mapping functions.
It's worth mentioning not all arbitrary stateful processing features are present in the PySpark API. It's not possible to initialize the state yet. If you are interested in this feature, you can track its progress or propose your implementation in the SPARK-40444.
Finally, take care of the JVM <=> PVM serialization loops. If your stateful function won't use all the columns available after reading the data, explicitly select the columns used in the mapping to reduce the serialization/deserialization overhead.
Example
To close this blog post, I'll leave you with an example of the arbitrary stateful processing in PySpark. The code reads the rate-micro-batch data source and keeps some time-based statistics for each group. It's available on my Github. Happy streaming!
Data Engineering Design Patterns

Looking for a book that defines and solves most common data engineering problems? I wrote
one on that topic! You can read it online
on the O'Reilly platform,
or get a print copy on Amazon.
I also help solve your data engineering problems contact@waitingforcode.com 📩
Read also about Arbitrary stateful processing in PySpark with applyInPandasWithState here:
Related blog posts:
- Spark Declarative Pipelines internals
- Spark Declarative Pipelines, going further
- Spark Declarative Pipelines 101
In the previous release #PySpark has got an interesting streaming feature -> the arbitrary stateful processing. It has a different API than the Scala version but is more adapted to the Python world.
— Bartosz Konieczny (@waitingforcode) September 29, 2023
More ? https://t.co/KfzgtIby32 pic.twitter.com/5PjoyDq0Dt

